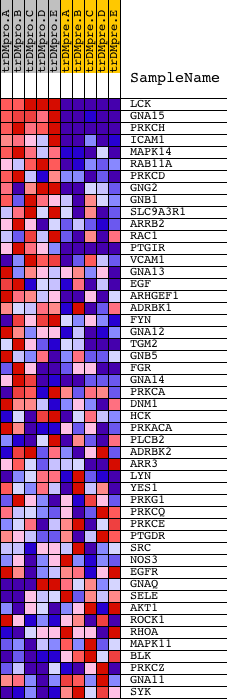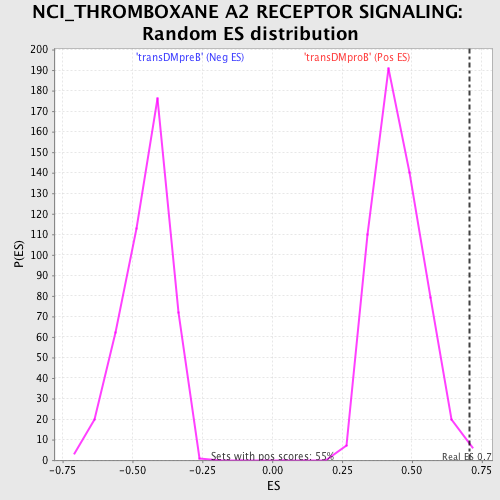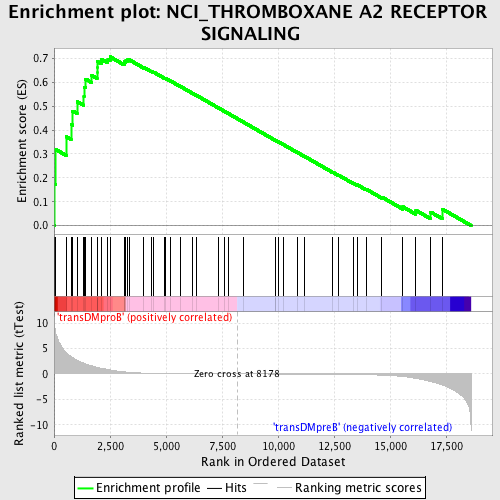
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | Set_04_transDMproB_versus_transDMpreB.phenotype_transDMproB_versus_transDMpreB.cls #transDMproB_versus_transDMpreB.phenotype_transDMproB_versus_transDMpreB.cls #transDMproB_versus_transDMpreB_repos |
| Phenotype | phenotype_transDMproB_versus_transDMpreB.cls#transDMproB_versus_transDMpreB_repos |
| Upregulated in class | transDMproB |
| GeneSet | NCI_THROMBOXANE A2 RECEPTOR SIGNALING |
| Enrichment Score (ES) | 0.7068897 |
| Normalized Enrichment Score (NES) | 1.5690044 |
| Nominal p-value | 0.0036166366 |
| FDR q-value | 0.31813186 |
| FWER p-Value | 0.804 |

| PROBE | DESCRIPTION (from dataset) | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|---|
| 1 | LCK | 15746 | 27 | 9.283 | 0.1731 | Yes | ||
| 2 | GNA15 | 19671 | 81 | 7.880 | 0.3185 | Yes | ||
| 3 | PRKCH | 21246 | 544 | 4.206 | 0.3727 | Yes | ||
| 4 | ICAM1 | 19545 | 760 | 3.333 | 0.4238 | Yes | ||
| 5 | MAPK14 | 23313 | 828 | 3.176 | 0.4799 | Yes | ||
| 6 | RAB11A | 7013 | 1027 | 2.668 | 0.5194 | Yes | ||
| 7 | PRKCD | 21897 | 1332 | 2.074 | 0.5421 | Yes | ||
| 8 | GNG2 | 4790 | 1339 | 2.062 | 0.5805 | Yes | ||
| 9 | GNB1 | 15967 | 1402 | 1.958 | 0.6140 | Yes | ||
| 10 | SLC9A3R1 | 11250 | 1679 | 1.595 | 0.6291 | Yes | ||
| 11 | ARRB2 | 20806 | 1919 | 1.273 | 0.6402 | Yes | ||
| 12 | RAC1 | 16302 | 1922 | 1.265 | 0.6639 | Yes | ||
| 13 | PTGIR | 18376 | 1926 | 1.262 | 0.6875 | Yes | ||
| 14 | VCAM1 | 5851 | 2122 | 1.045 | 0.6966 | Yes | ||
| 15 | GNA13 | 20617 | 2402 | 0.850 | 0.6976 | Yes | ||
| 16 | EGF | 15169 | 2502 | 0.778 | 0.7069 | Yes | ||
| 17 | ARHGEF1 | 1952 18343 3996 | 3132 | 0.392 | 0.6804 | No | ||
| 18 | ADRBK1 | 23960 | 3152 | 0.380 | 0.6865 | No | ||
| 19 | FYN | 3375 3395 20052 | 3206 | 0.359 | 0.6904 | No | ||
| 20 | GNA12 | 9023 | 3258 | 0.338 | 0.6940 | No | ||
| 21 | TGM2 | 2657 | 3344 | 0.305 | 0.6952 | No | ||
| 22 | GNB5 | 2985 9029 3153 | 4007 | 0.151 | 0.6624 | No | ||
| 23 | FGR | 4723 | 4353 | 0.105 | 0.6458 | No | ||
| 24 | GNA14 | 23907 | 4444 | 0.097 | 0.6428 | No | ||
| 25 | PRKCA | 20174 | 4927 | 0.066 | 0.6180 | No | ||
| 26 | DNM1 | 2777 8857 | 4970 | 0.064 | 0.6170 | No | ||
| 27 | HCK | 14787 | 5214 | 0.056 | 0.6050 | No | ||
| 28 | PRKACA | 18549 3844 | 5625 | 0.043 | 0.5837 | No | ||
| 29 | PLCB2 | 5262 | 6166 | 0.031 | 0.5552 | No | ||
| 30 | ADRBK2 | 11499 | 6353 | 0.027 | 0.5457 | No | ||
| 31 | ARR3 | 24282 9305 | 7321 | 0.012 | 0.4938 | No | ||
| 32 | LYN | 16281 | 7605 | 0.008 | 0.4787 | No | ||
| 33 | YES1 | 5930 | 7788 | 0.005 | 0.4690 | No | ||
| 34 | PRKG1 | 5289 | 8458 | -0.004 | 0.4331 | No | ||
| 35 | PRKCQ | 2873 2831 | 9879 | -0.022 | 0.3570 | No | ||
| 36 | PRKCE | 9575 | 9886 | -0.022 | 0.3571 | No | ||
| 37 | PTGDR | 21866 | 10024 | -0.023 | 0.3501 | No | ||
| 38 | SRC | 5507 | 10226 | -0.026 | 0.3398 | No | ||
| 39 | NOS3 | 16906 885 | 10879 | -0.037 | 0.3054 | No | ||
| 40 | EGFR | 1329 20944 | 11172 | -0.042 | 0.2905 | No | ||
| 41 | GNAQ | 4786 23909 3685 | 12422 | -0.073 | 0.2246 | No | ||
| 42 | SELE | 14074 | 12690 | -0.083 | 0.2117 | No | ||
| 43 | AKT1 | 8568 | 13376 | -0.114 | 0.1770 | No | ||
| 44 | ROCK1 | 5386 | 13538 | -0.125 | 0.1707 | No | ||
| 45 | RHOA | 8624 4409 4410 | 13936 | -0.158 | 0.1522 | No | ||
| 46 | MAPK11 | 2264 9618 22163 | 14624 | -0.244 | 0.1198 | No | ||
| 47 | BLK | 3205 21791 | 15569 | -0.514 | 0.0787 | No | ||
| 48 | PRKCZ | 5260 | 16150 | -0.885 | 0.0641 | No | ||
| 49 | GNA11 | 3300 3345 19670 | 16806 | -1.493 | 0.0569 | No | ||
| 50 | SYK | 21636 | 17335 | -2.157 | 0.0690 | No |